Machine learning for classifying chronic kidney disease and predicting creatinine levels using at-home measurements
We separated the features into the three sets, as outlined in Table 1, based on the tools required for measurement. At-home features are known by a patient or are easily measurable, like age or blood pressure. Monitoring features are obtainable from regular check-ups, such as red blood cell count, and laboratory features are from blood and urine tests targeted towards CKD, such as urine albumin.
We have classified CKD and predicted creatinine levels on the three groups of features. In the classification case, we classified whether a person has CKD or not, disregarding stages. A patient’s GFR, and thus stage, can be estimated using the predicted creatinine level as an input into the CKD-EPI equation8. We carried out our experiments using both ANN and RF. Using the monitoring and laboratory feature sets, both ANN and RF had near perfect classification accuracy, and RF performed better than ANN using the at-home features. This is further highlighted by the ROC curves and AUCs obtained (Fig. 2). With monitoring and laboratory feature sets, ANN and RF both had AUCs of essentially 1. With the at-home feature set RF performed slightly better than ANN with a higher AUC, and a better ROC with a lower variance. Similarly, for creatinine regression both ANN and RF have comparable results using the monitoring and laboratory features, and RF performed better using at-home features. We observed some distinct performance differences between ANN and RF, particularly with at-home features. ANN achieved an average accuracy of 82.9%, a TPR of 92.0%, and a TNR of 67.9%, while RF performs better with a 92.5% accuracy, 90.0% TPR, and notably higher 95.8% TNR, making RF more suitable for at-home CKD classification due to its reduced false negative rate. For both monitoring and laboratory features, however, ANN and RF perform comparably, with accuracy, TPR, and TNR values all over 98%, showing high effectiveness with more comprehensive data. The ROC curves align with these results, with RF slightly outperforming ANN for at-home features (AUC of 0.965 vs. 0.936), while both methods show near-perfect AUC values with monitoring and laboratory features. For creatinine regression, RF achieves a higher R2 with at-home features (0.381), though this remains suboptimal, indicating limited predictive power without detailed clinical data. Overall, RF demonstrates a modest advantage with at-home features, while both models perform equally well when more extensive data is available. The advantage of RF over ANN when using at-home features is somewhat expected, as a majority of these features are categorical, and RF naturally handles categorical data more effectively than ANN due to the structure of decision trees. We expect the performance of ANN to improve with a more complex architecture, but this would require much more additional data.
Exploiting the nature of RF algorithms, we extracted the most important features in the classification and regression (Fig. 3). Hypertension and diabetes mellitus were the two most important features of the at-home set for both classification and regression as measured by the mean entropy importance. We had less agreement of the most impactful features between the classification and regression on the monitoring and laboratory feature sets. For monitoring features, blood urea and sodium are highly important for regression, but are less important for CKD classification. Red blood cell ranks second in classification, but does not appear in the top 10 for regression. Similarly, diabetes mellitus is among the top five in classification, but is much less significant for regression; hypertension remains important for both tasks. In the laboratory feature set, blood urea emerges as crucial for regression, while, again, red blood cell ranks fourth in classification, but is absent from the top 10 for regression. Hypertension is fifth in classification, but does not appear in the top 10 for regression. Hemoglobin was the most important feature for both monitoring and laboratory sets in the classification task, and both red blood cell count and hypertension were in the top five for both sets. When predicting creatinine, blood urea was by far the most important feature within the monitoring and laboratory sets, with hemoglobin, red blood cell count, and sodium being in the top five for both sets.
In our study, we leverage a dataset that has been examined in prior works18,24,25,26. The common thread among these studies, like one of our own, is the focus on CKD classification using all available features. Across these studies and our own (with monitoring or laboratory features), we consistently observe near-perfect accuracy in CKD classification, highlighting the robustness of machine learning methods. Presently, CKD diagnoses require laboratory measurements at least three months apart6, hence, machine learning could reduce wait time for a diagnosis and treatment plan.
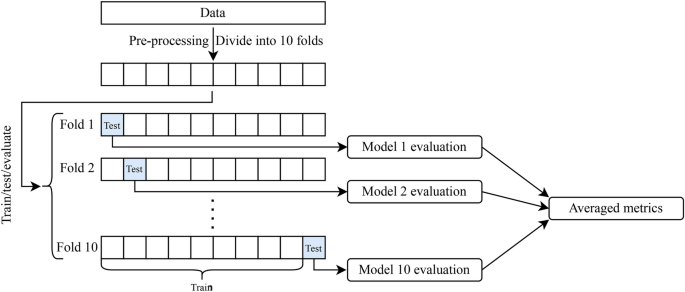
Both Khalid et al.24 and Almansour et al.25 explore subsets of this dataset by either using only numerical features or by examining the performance with a reduced number of features, respectively. Specifically, Khalid et al.24 applied a 10-fold cross-validation approach using 14 features, including serum creatinine, and tested models such as gradient boosting, Gaussian Naive Bayes, decision tree, RF, and a hybrid model. They only report accuracy, and achieved an accuracy of 98% for RF and 100% for their hybrid model. Similarly, Almansour et al.25 employed support vector machine and ANN with 10-fold cross-validation, focusing on “best” feature subsets. They reported accuracies of 97.75% on the full feature set for both models and, notably higher accuracies of 98.5% and 98% using the best 12 features for support vector machine and ANN, respectively. Our results match or surpass these findings, as we achieve higher accuracy with both ANN and RF using the monitoring (98.7% for both) or laboratory (99.2% and 99.5%, respectively) feature sets. Additionally, our study contributes a novel perspective by categorizing features into at-home, monitoring, and laboratory subsets. Qin et al.18 tested multiple models, including logistic regression, RF, support vector machine, k-nearest neighbours, Naive Bayes, ANN, and an integrated model. They achieved over 99% accuracy, TPR, and TNR for logistic regression, RF, and the integrated model. Qin et al.18 identified key features such as specific gravity, hemoglobin, serum creatinine, albumin, packed cell volume, red blood cell count, hypertension, and diabetes mellitus as significant for CKD classification. Our findings align closely with theirs, underscoring the value of these attributes in CKD diagnosis. Pal26 employed support vector machine, RF, and ANN for binary classification using categorical, non-categorical, and combined feature sets, including serum creatinine. They reported accuracies of 88%, 92%, and 80% for support vector machine, RF, and ANN, respectively, along with TPRs of 61%, 55%, and 55% and AUC values of 0.77, 0.76, and 0.70, respectively. In contrast, our study achieves near-perfect results using the laboratory or monitoring feature sets (while excluding serum creatinine), with both ANN and RF models achieving a mean accuracy exceeding 99% and AUC values of 0.999 and 1.00, respectively. Unlike previous studies that focus on subsets of “best” features, we emphasize grouping data that is typically collected together, which provides a practical framework for CKD prediction based on context-specific data availability. Our approach enhances the interpretability of our models and provides insights into the relevance of features for different aspects of CKD prediction.
We employed a similar approach to Wang et al.14 by predicting creatinine levels directly. Their ensemble method achieved an R2 score of 0.5590 using a different dataset, and they emphasize the significance of hemoglobin. In our findings, hemoglobin was in the top three features for both monitoring and laboratory features for creatinine prediction. We note the data used in Ref.14 did not contain blood urea, which we found to be the most significant predictor.
A comprehensive overview of machine learning techniques for CKD classification can be found in Ref.11, where a wide range of methods are assessed. Notably, the best-performing models in their tabulation achieve accuracies of 98% or higher. Sanmarchi et al.12 conducted a review encompassing 68 relevant articles on CKD prediction, diagnosis, and treatment using a wide range of machine learning methods. Their findings emphasize the importance of attributes such as blood pressure, hemoglobin, sodium, albumin, pus cell, red blood cell count, and diabetes mellitus for CKD prognosis and diagnosis. These highlighted features align with our own feature importance observations, accentuating their relevance in the context of CKD assessment.
In summary, our study builds upon a dataset examined in previous research and offers a unique perspective by categorizing features into context dependent subsets. Our findings, including the importance of specific attributes and the success of machine learning methods in CKD classification, corroborate and extend upon existing literature, contributing to a better understanding of CKD detection and prediction.
Our study has some limitations however, that warrant consideration. Firstly, while the CKD and not CKD labels in the dataset were assigned by nephrologists using patient history, symptoms, and blood and urine tests, it is not clear precisely what criteria were used to determine the labels. Moreover, the absence of stage-specific information for patients with CKD in our dataset poses a challenge. Additionally, our models’ ability to detect early-stage CKD may be limited, as we primarily focused on classifying CKD in general, not the stage. Having labelled data with explicit CKD stages would enable a more nuanced analysis and classification of disease progression. Presently, our creatinine regression results provide a proxy for stage, but this transition from creatinine to stage introduces additional uncertainty, given the equations involved are empirical in nature. These limitations highlight the need for more comprehensive and stage-specific datasets to further improve the accuracy and clinical relevance of CKD detection and classification models. We also acknowledge the necessity for larger and more comprehensive datasets to enhance the precision and applicability of our models. Additionally, incorporating a more diverse population is crucial to ensure a model’s robustness across various demographics and regions. These improvements would lead to more accurate and generalizable predictions in real-world clinical settings.
link