Cockayne syndrome mice reflect human kidney disease and are defective in de novo NAD biosynthesis
Animals
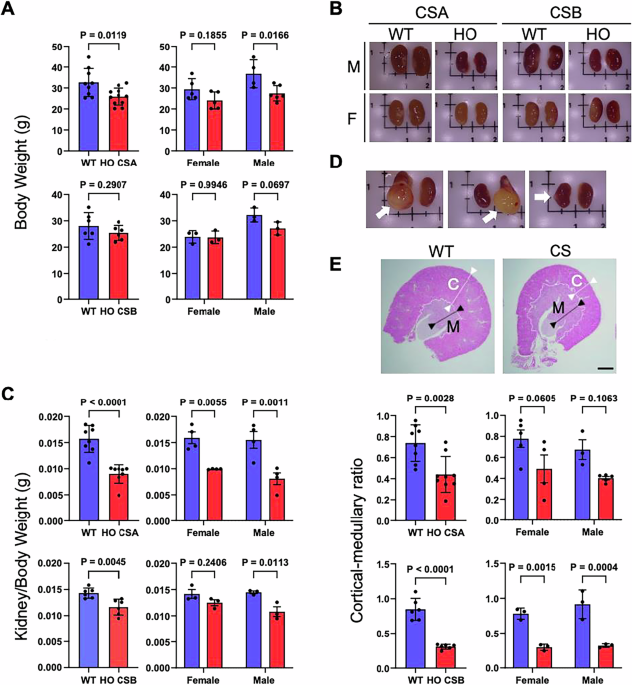
Mouse models of CSA−/− (knockout mice), CSBm/m (mice carrying a premature stop codon in exon 5 to mimic the K337 stop truncation mutation from a CS patient), and wild-type (WT) on a C57BL/6 J background in the age range of 45–68 weeks were used for all the experiments, unless otherwise specified.
Antibodies
The antibodies used were as follows: CSA (Abcam, ab137033), CSB (Abcam, ab96089), QPRT (Biorybt, orb317756), ATF3 (Abcam, ab207434), β Actin (SantaCruz, sc47778), GAPDH (Cell Signallng Technology, 2118), Havcr-1 (ThermoFischer scientific, PA5-98302).
Cell Culture
HK-2 (Human kidney proximal tubular epithelial cell line (ATCC CRL-2190)) cells were cultured in Keratinocyte serum-free medium (K-SFM) containing bovine pituitary extract (BPE), human recombinant epidermal growth factor (EGF) in a humidified chamber under 5% CO2 at 37 °C.
siRNA knockdown
siRNAs for CSA (SR319723A; SR319723C) and CSB (SR320072A; SR320072B) and non-targeted siRNA (SR3005) were purchased from Origene (Rockville, MD, USA). siRNA was diluted with K-SFM to a final concentration of 50 nM, mixed with JetPrime (Polyplus transfection, Illkirch-graffenstaden, France), incubated for 15 min at room temperature, and transfected into target cells according to the manufacturer’s instructions. Twenty‐four hours after transfection, the media was replaced. Cells were harvested 72 h after transfection and used for the experiments.
NR supplementation
72 h post siRNA transfection, HK-2 cells were supplemented with 1 mM NR (Chromadex, CA, USA) for 24 h. Following NR treatment, the cells were harvested and used for further experiments.
Immunoblotting
Cells were harvested, washed with PBS, and lysed in 1X RIPA buffer (ThermoFisher scientific, Waltham, MA, USA) with protease inhibitor cocktail (Cell Signaling Technology, Danvers, MA, USA). Protein concentrations were determined by a BCA (ThermoFisher scientific) assay. Equal concentrations of proteins were loaded on Mini-PROTEAN® precast gels (Bio-Rad, Hercules, CA, USA) for SDS-PAGE. Proteins were transferred on a PVDF membrane (0.45 μM pore size) using a Mini Trans-Blot® Cell transfer system (Bio-Rad). After blocking the membrane in 3% nonfat milk in TBS-T (Tris buffered saline with 0.1% Tween-20), it was incubated overnight with primary antibodies at 4°C, followed by anti-rabbit HRP-conjugate IgG or anti-mouse HRP-conjugate IgG. Immunoblot signals were detected using an ECL detection kit (ThermoFisher scientific). Immunoblots were imaged and quantitated using a Bio-Rad Chemidoc platform.
NAD+ quantification
Kidneys were dissected in cold PBS and placed in NADH/NAD Extraction Buffer (ab65348, abcam, Boston, MA, USA). Following homogenization with a micro pestle, an NAD/NADH Assay Kit (ab65348, abcam) was used to quantify NAD+ and NADH levels.
HK-2 cells were harvested after siRNA transfection, washed with PBS, and lysed in NADH/NAD Extraction Buffer (ab65348, abcam). NAD+ and NADH levels were quantified according to the manufacturer’s instructions. Data were normalized to scrambled siRNA treated (SCR) HK-2 cells.
Quantitative real-time PCR (qPCR)
Total RNA was extracted from cells using a Nucleospin Triprep kit (Macherey-Nagel, Düren, Nordrhein-Westfalen, Germany) or Purelink RNA isolation kit (Invitrogen, CA, USA). One microgram of total RNA was reverse transcribed using the iScript™ cDNA Synthesis Kit (Bio-Rad). qPCR was performed using the DyNAmo HS SYBR Green qPCR Kit (F-410L, ThermoFisher scientific) with the CFX Connect Real-time PCR Detection System (Bio-Rad). Ct values were normalized to values for GAPDH for HK-2 cells while Actin was used for mice. Primer sequences used are listed in Supplementary Table 1. Data are reported as mean ± standard deviation.
RNA sequencing
RNA from the mice were isolated using RNeasy (Macherey-Nagel). Library construction and sequencing were performed by Novogene (Sacramento, CA, USA). The samples were run on the NovaSeq 6000. The samples have >95% bases with Q20 and above. Samples were aligned to the reference mouse genome (mm10) and junctions using HISAT2 (v2.0.5) software. Differential expression analysis of the groups (two biological replicates per condition) were performed using the DESeq2 R package (1.20.0). The resulting P-values were adjusted using the Benjamini and Hochberg’s approach for controlling the false discovery rate (FDR). Genes with an adjusted P value ≤ 0.05 found by DESeq2 were assigned as differentially expressed with foldchange of 2 and 0.5.
Gene Ontology (GO) enrichment analysis of differentially expressed genes was implemented by the ClusterProfiler R package, in which gene length bias was corrected. GO terms with corrected P value less than 0.05 were considered significantly enriched by differential expressed genes. ClusterProfiler R package was used to test the statistical enrichment of differential expression genes in KEGG pathways.
GEO accession number is GEO # GSE246444.
Biological assessment
Whole blood samples from the mice were collected at sacrifice by cardiac puncture. Blood was allowed to stand for 30 min and centrifuged at 2000 × g for 15 min. Serum was transferred to a new tube and different biochemical parameters were checked. Urine was collected from the mice by holding the mice over the collection tube, until sufficient amounts were collected for biochemical analysis. Biochemical analysis for Serum and urine was done by IDEXX Laboratories, Inc (Westbrooke, Maine, USA). Urine chemistry values were normalized to each animal’s respective urine creatinine level.
Detection of kidney toxicity markers
Serum and urine collected from the mice were analyzed using a FirePlex® kidney toxicity immunoassay (Abcam, ab235661), to detect the levels of clusterin, cystatin C, havcr1, lipocalin-2, and osteopontin as per the manufacturer’s protocol.
Chromatin immunoprecipitation (ChIP)
ChIP was performed as described [20], with slight modifications. Briefly, HK-2 cells were incubated in 1% formaldehyde PBS for 10 min at room temperature. The solution was brought to a final concentration of 0.125 M glycine, and cells were harvested by centrifugation, washed three times in cold PBS, then incubated in buffer A (5 mM PIPES (pH 8.0), 85 mM KCl, 0.5% NP-40, and protease inhibitor cocktail (GenDEPOT, Katy, TX, USA)). Cell extracts were centrifuged, and pelleted material was resuspended in buffer B (100 mM Tris-Cl (pH 8.1), 1% sodium dodecyl sulfate (SDS), 10 mM EDTA, and protease inhibitor cocktail). Chromatin was sheared using an S-450 sonicator (Branson, Danbury, CT, USA.). An aliquot containing 500 μg DNA was diluted 10-fold in IP buffer (0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-Cl (pH 8.1), 167 mM NaCl, and protease inhibitor cocktail) and incubated with primary antibody for ATF3 (abcam ab207434) overnight at 4°C. Samples were incubated 30 min at 4 °C with protein A/G-linked magnetic beads. Beads were washed sequentially with TSE150 (1% Triton X-100,0.1% SDS, 2 mM EDTA, 20 mM Tris-Cl (pH 8.1), and 150 mM NaCl), TSE500 (1% Triton X-100, 0.1% SDS, 2 mM EDTA, 20 mM Tris-Cl (pH 8.1), and 500 mM NaCl), and Buffer III (0.25 M LiCl, 1% NP-40, 1% sodium deoxycholate, 1 mM EDTA, and 10 mM Tris-Cl (pH 8.1)) and then washed twice with TE (pH 8.0) for 10 min. Chromatin was eluted with elution buffer (1% SDS and 0.1 M NaHCO3 (pH 8.0)) and incubated overnight at 65°C in 200 mM NaCl to reverse cross-linking. Aliquots (500 μL) were incubated at 50°C after addition of 10 μL 0.5 M EDTA, 20 μL 1 M Tris (pH 6.5) and 4 μL Proteinase K (20 mg mL−1), extracted sequentially with phenol/chloroform/isoamyl alcohol. Nucleic acids were pelleted by centrifugation for 30 min at 4°C after addition of 1 μL of 20 mg mL−1 glycogen, 20 μL of 5 M NaCl and 500 μL of isopropanol. Pellets were washed with 70% ethanol, dried and resuspended in nuclease-free water.
qPCR of ChIP products
ChIP products were subject to qPCR using the DyNAmo HS SYBR Green qPCR Kit (F-410L, ThermoFisher scientific) with the CFX Connect Real-time PCR Detection System (Bio-Rad). Experimental values were normalized to the values of 1% input chromatin. Concentrations were estimated using the 2−ΔΔCT calculation method. The sequences of the primers are listed in Supplementary Table 1.
Histological analysis
Mice were anesthetized deeply with isoflurane and then perfused with 1X phosphate buffered saline (PBS) through the heart. Kidneys were removed, fixed in 4% paraformaldehyde (PFA) for 24 h at 4 °C, and then washed with PBS. Tissue preparation, mounting, and staining was done by Histoserv (Germantown, MD, USA). Kidneys were trimmed, processed, and embedded in paraffin blocks. Formalin-fixed, paraffin-embedded (FFPE) blocks were cut into 5 µm thick sections and placed on Superfrost slides. To perform immunohistochemistry, slides were deparaffinized using xylene and rehydrated using graded concentrations of ethanol and water. Heat-mediated antigen retrieval was then performed using a citrate-based buffer. Slides were then incubated with the primary antibody, either QPRT (orb317756, Biorybt, Cambridge, UK) (1:1000) or Havcr-1 (PA5-98302, ThermoFisher scientific) (1:1600) followed by a goat-anti-rabbit-HRP-conjugated secondary. Detection was then developed with DAB and counterstained with hematoxylin. Slides were subsequently dehydrated, cleared, mounted with a permanent mounting medium, and scanned using a Leica AT2 system at 0.25 µm/px. Image analysis and quantification for Havcr1 was using Qupath [21], while QPRT images were analyzed and quantitated by ImageJ [22].
Sections (4 μm thick) cut from 10% formalin-fixed, paraffin-embedded kidney samples were used for periodic acid-Schiff (PAS) staining or picrosirius red (PSR) staining from Histoserv. Quantifications were performed in a masked manner. Using coronal sections of the kidney, 30 consecutive glomeruli per mouse, with 3–5 mice per group, were examined for evaluation of glomerular mesangial expansion. The index of mesangial expansion was scored based on the ratio of mesangial area to glomerular tuft area. The mesangial area was determined by assessment of PAS-positive and nucleus-free areas in the mesangium using the ScanScope image analyzer (Aperio Technologies, Vista, CA). Using color deconvolution algorithm on ScanScope image analyzer, the quantification of PSR staining was expressed as a percentage of the red positive area on the total cortical surface.
TUNEL assay
A TUNEL assay was performed using the Click-iT™ Plus TUNEL Assay for In Situ Apoptosis Detection kit (Invitrogen, CA, USA). In brief, HK-2 cells were transfected with CSA/B siRNAs. After 72 h of transfection, cells were supplemented with/without 1 mM NR for 24 h. Following NR treatment, cells were washed three times with PBS, fixed with 4% formaldehyde for 30 min, and permeabilized with 0.5% Triton X-100 for 10 min. Then, the TUNEL probe was incubated with the cells for 4 h at room temperature. After staining with Hoechst to label the nucleus for 3 min at room temperature, the samples were observed and analyzed under an inverted microscope (Zeiss, Observer Z.1).
TUNEL staining for kidney sections was performed using a TUNEL assay kit from abcam (ab20638), following the manufacturer’s protocol.
p57 immunohistochemistry
Paraffin-embedded kidney section slides were deparaffinized and rehydrated using graded concentrations of ethanol and water. Heat-mediated antigen retrieval was then performed using an EDTA-based buffer. After retrieval, slides were then incubated with the p57 primary antibody (ab75947, abcam, Waltham, MA) (1:100) in antibody diluent (BSB 0114, BioSB Inc., Santa Barbara, CA) followed by a goat-anti-rabbit-HRP-conjugated secondary antibody (AF835, R&D system, Minneapolis, MN). Detection was then developed with an AEC substrate (SK-4205, Vector Laboratories, Newark, CA) and counterstained with hematoxylin. Slides were mounted with Vecta Mount AQ mounting medium (Vector Lab, H-5501) and scanned using a MoticEasyscan scanner (Motic, Vancouver, Canada). The slides were stripped and re-stained with the periodic acid Schiff stain reagents (87007, Richard-Allan Scientific, Kalamazoo, MI). Images were re-collected by scanning p57, and periodic acid-Schiff-stained images for each slide, which were then fused into a single image by HALO software (Indica Labs, Albuquerque, NM). The average number of podocytes per glomerulus was calculated by manually counting 50 glomeruli for each case section.
Glomerulus quantification from digital histology
Glomeruli were manually annotated using the Aperio ImageScope digital histology viewer [v12.3.3, Leica Biosystems]. All subsequent computational image analysis was completed in MATLAB [vR2024b]. Whole-slide glomerulus masks were computed from xml annotation files, and glomerulus areas were computed using the regionprops function. Mesangium was segmented from each glomerulus using color deconvolution with HPAS stain vectors extracted using the Fiji ImageJ Color Deconvolution plug-in. The percentage mesangial area was computed as the sum of PAS+ pixels indexed to glomerular area.
Glomerular area feature mapping
The areas of glomeruli from all wild-type kidneys were pooled, and a histogram was constructed using a sequential color heatmap. In the color heatmap, low glomerulus areas were represented by dark red, while the highest glomerulus areas were represented by dark blue. The glomeruli from all KO kidneys were then assigned to the corresponding histogram bin based on their glomerular area. Glomeruli in whole-slide image segmentation masks were re-colored according to the histogram bin that they were assigned, producing images where glomeruli are colored based on area relative to the global distribution of glomerulus areas across the dataset. Re-colored glomerulus segmentation masks were superimposed on whole-slide H&E images to visualize the underlying glomerular histologic presentation. All image processing was completed in MATLAB.
Tubule quantification from digital histology
ARTSA, a desktop application for renal tubule quantification from whole-slide histopathology, was used to automatically segment and quantify tubules and tubule nuclei [23, 24]. Morphological features extracted from renal tubules included area, perimeter, eccentricity, solidity, and percentage tubule (total tubule area per kidney indexed to total kidney area). Percentage nuclear area per tubule was computed as the total nuclear area per tubule indexed to tubule area. Textural features extracted from tubule nuclei included first and second-order pixel statistics. First order nuclear pixel statistics included minimum, maximum, range, mean, variance, standard deviation, median, interquartile range, 10th percentile, 90th percentile, mean absolute deviation, root mean absolute deviation, root mean squared, skewness, kurtosis, uniformity, and entropy. Second order statistics included gray-level co-occurrence matrix (GLCM) contrast, correlation, energy, and homogeneity.
Statistical analysis
GraphPad Prism (GraphPad Software, Inc.) was used to perform the statistical analysis. A two-tailed t test was used to compare individual group. A two-way ANOVA was used to compare multiple samples within a single group.
For all computed histological image features, normality was assessed with the Anderson Darling test, and variance was assessed with Bartlett’s (parametric) or Levene’s (non-parametric) test. For comparisons among groups, ANOVA (parametric) or Kruskal-Wallis (non-parametric) tests were completed, with corresponding post-hoc tests (Tukey’s and Dunn’s tests, respectively).
link