Analyzing longitudinal trait trajectories using GWAS identifies genetic variants for kidney function decline

UKB eGFR-trajectories exhibit an approximately linear decline of −1 mL/min/1.73 m2/year
We analyzed unrelated European-ancestry UKB individuals without acute kidney injury (AKI) or nephrectomy, excluding eGFR assessments after onset of dialysis, kidney transplant, or end-stage kidney disease (ESKD) (“Methods” section). Our analyzed UKB data consisted of 149,263 individuals with ≥2 eGFR assessments per person (“UKB 150K”; median follow-up time = 8.4 years; m = 1,321,370 eGFR assessments) or 348,275 individuals with ≥1 eGFR assessment (“UKB 350K”; m = 1,520,382; Supplementary Fig. 1). UKB 350K was similar to 150K regarding participant characteristics: 54% women, 1.2% CKD at baseline and 4.6% at any timepoint (eGFR < 60 mL/min/1.73 m2), baseline age 35–78 years, median baseline eGFR = 97 mL/min/1.73 m2 (Table 1). We used UK10K/HRC-imputed allele dosages of 11.3 million single-nucleotide polymorphisms (SNPs) and selected 595 variants known for association with cross-sectional eGFR23 (“Methods” section).
Before evaluating genetic variants, we explored a potentially non-linear relationship of eGFR with time and age, observing approximate linearity and negligible difference by sex (Supplementary Fig. 2a–c). This was more challenging for individuals with CKD, primarily due to regression-to-the-mean effects at the start of trajectories and sparse data at their end (Supplementary Fig. 2d). Assuming linearity, mean annual eGFR-decline was comparable across approaches (−0.88 to −1.08 mL/min/1.73 m2/year), with high variability of person-specific slopes (standard deviation 0.66–0.95 mL/min/1.73 m2/year, Supplementary Table 1 and Supplementary Note 1).
LMM age model RI&RS is a powerful approach with unbiased genetic effect estimates
We considered seven approaches for genetic association analysis with eGFR-decline (Supplementary Table 2, “Methods” section Eqs. (1–4)): in data of individuals with ≥2 assessments over time, (i) difference model, (ii–v) four one-stage LMMs (time model RI&RS, age model RI&RS, age model RI&RS uncorrelated, age model RI-only), (vi) an LMM-based two-stage approach (BLUPs&LinReg); in data adding singletons (i.e., individuals with =1 assessment), (vii) age model RI&RS.
We compared these approaches in simulated data using various scenarios (simulation parameters corresponding to: eGFR-trajectories as in UKB 350K, ~50% singletons; eGFR-trajectories in an external cohort study, KORA-424, ~20% singletons; trajectories of another trait, body mass index, BMI, in KORA-4; “Methods” section, Supplementary Table 3). We found the following (Table 2 and Supplementary Table 4): (i) type I error was inflated for age model RI-only and age model RI&RS uncorrelated, indicating insufficient accounting for person-specific slope variability. (ii) Power was better for one-stage LMMs compared to difference model, but BLUPs&LinReg was the most powerful. When adding singletons, not possible with difference model or BLUPs&LinReg, the age model RI&RS became nearly as powerful as BLUPs&LinReg in the UKB-based scenario. (iii) Biased effect estimates were observed for BLUPs&LinReg in all scenarios (11%–38% shrinkage), in line with the bias-variance trade-off known from regularization25 (Supplementary Note 2), while estimates from age model RI&RS were unbiased.
Empirical data (UKB 150K, or 350K when adding singletons) corroborated simulation findings regarding type I error (no control by age model RI-only and RI&RS uncorrelated, Supplementary Fig. 3), power (best for BLUPs&LinReg and age model RI&RS in UKB 350 K), and bias (BLUPs&LinReg: 38.5% shrinkage; Table 2, Supplementary Note 2, Supplementary Fig. 4, Supplementary Data 1).
Altogether, among approaches with type I error control, BLUPs&LinReg showed the best power, but biased effect estimates. When jointly aiming for good power and unbiased effect estimates, the LMM age model RI&RS was preferable, particularly in the UKB 350K dataset. We thus used the LMM age model RI&RS in UKB 350K in the following.
Twelve genetic variants across ten loci identified for association with eGFR-decline
Due to our hypothesis that genetics of eGFR-decline is a subset of genetics of cross-sectional eGFR, we first focused on the 595 variants known for cross-sectional eGFR-association23 and tested these for association with eGFR-decline (“595-search”, LMM age model RI&RS in UKB 350 K). We identified 12 variants (Pdecline < 0.05/595 = 8.4 × 10−5, 6 with Pdecline < 5 × 10−8, Fig. 2a and Table 3): (i) 7 variants known for eGFR-decline10 (near/in UMOD/PDILT (2), TPPP, C15orf54, FGF5, OVOL1, and PRKAG2) and (ii) 5 variants novel for eGFR-decline: 1 independent third UMOD/PDILT variant and 4 novel loci (near SDCCAG8, RRAGD, GGT7, PRAG1). We raised the number of variants with Pdecline < 5 × 10−8 from two (UMOD/PDILT) to six (four loci, adding loci around TPPP, C15orf54, SDCCAG8; Table 3). Results were robust upon various sensitivity analyses (Supplementary Fig. 5 and “Methods” section).
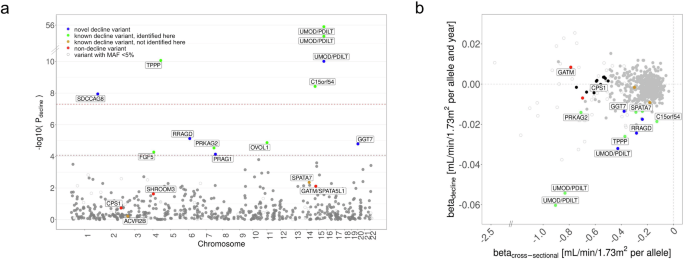
We selected 595 SNPs previously reported for association with eGFR in cross-sectional data23 and tested them for association with eGFR-decline using the one-stage LMM age model RI&RS 350K (UKB 350K; n = 348,275, m = 1,520,382). a Shown are P values (Pdecline) versus chromosomal position. We identified 12 variants (10 loci) for eGFR-decline at Bonferroni(595)-corrected significance (Pdecline < 0.05/595 = 8.4 × 10−5, brown dashed horizontal line; including 6 with Pdecline < 5 × 10−8, red dashed horizontal line), consisting of 5 novel and 7 known variants for eGFR-decline10 (blue or green, respectively). Also color-coded are two variants known for eGFR-decline not identified here (orange) and three variants known for not being associated with eGFR-decline (red)10. Variants with small minor allele frequency (MAF < 5%) are shown as circles. b Shown are genetic effect sizes for eGFR-decline (βdecline from LMM age model RI&RS 350K) versus effect sizes for association with eGFR cross-sectionally (βcross-sectional: eGFR~sex, age, SNP, PCs; eGFR from UKB baseline study-center assessment, n = 341,073). Color and symbol codes are as in (a), additionally highlighting 11 stable-effect variants (black; Pmain < 5 × 10−8, |βmain| > 0.50 mL/min/1.73 m2/allele; Pdecline ≥ 0.1; |βdecline| < 0.005 and SEdecline < 0.005 mL/min/1.73 m2/allele and year) that include the CPS1 variant (rs1047891; red in (a)). Effect allele was the cross-sectionally eGFR-lowering allele (unconditioned analyses in EUR23). The exact numerical values are provided in Supplementary Data 2.
The five novel variants were detected with a similar number of individuals as in previous work10 (n ~ 350,000; CKDGen, difference model) due to the age model, not with the difference model in UKB or CKDGen or due to different multiple testing burdens (Table 3 and Supplementary Data 1).
Among the nine variants previously identified for eGFR-decline10, seven were identified here (Pdecline < 0.05/595), one additional variant had Pdecline = 5.1 × 10−3 (directionally consistent; Supplementary Table 5). We also confirmed variants near CPS1, SHROOM3, and GATM as not associated with eGFR-decline (Pdecline ≥ 0.05, Supplementary Table 5).
Validation in external data
We obtained support in independent longitudinal data: in three population-based cohort studies from Germany, we had previously reported an approximate linear relationship of eGFR over age26 (KORA-3: n = 2933, m = 3749; KORA-4: n = 3752, m = 9644; AugUR: n = 2397, m = 3442). Baseline age was 35–84, 25–74, or 70–95 years with ~20 years (KORAs) or ~9 years of follow-up (AugUR). The %CKD was higher in these studies than in UKB: %CKD at baseline (eGFR < 60 mL/min/1.73 m2) was 5.6%, 1.5%, and 21.5%, respectively, and %CKD at any timepoint was 6.7%, 8.2%, and 26.1%. The 12-variant polygenic score in combined KORA&AugUR data was significantly associated with eGFR-decline (Pdecline = 0.013; age model RI&RS, “Methods” section).
Decline-associated variants have little effect on eGFR for 40-year-old individuals and large effects on 70-year-old individuals in contrast to 11 stable-effect variants
When comparing directionality and size of variants’ effects on eGFR-decline with effects on cross-sectional eGFR (UKB study-center baseline, n = 341,073, aged 39–72 years), we found the 12 decline-accelerating alleles to coincide with cross-sectionally eGFR-lowering alleles (Fig. 2b, blue and green dots; Supplementary Data 2). One “bad” allele lowered average eGFR by −0.012 to −0.060 mL/min/1.73 m2/year compared to cross-sectional effects of −0.13 to −0.90 mL/min/1.73 m2 (Supplementary Data 3). We also observed variants with large cross-sectional effects that had no association with eGFR-decline (e.g., CPS1 variant).
We extracted variants with large main effect on eGFR-levels and no association with eGFR-decline (Pmain < 5 × 10−8, |βmain| > 0.50 mL/min/1.73 m2 per allele, Pdecline ≥ 0.1, |βdecline| < 0.005 and SEdecline < 0.005 mL/min/1.73 m2 per allele and year), yielding 11 “stable-effect” variants (including CPS1; Supplementary Data 3). Their main effects, reflecting genetic effects on eGFR for 50-year-old individuals due to age-centering, were similar to cross-sectional effects (βcross-sectional = −0.50 to −0.74 mL/min/1.73 m2; Fig. 2b, black dots).
We visualized the 12 + 11 SNP associations on eGFR-levels over age (βmain + (age-50)*βdecline): the 12 decline-associated variants showed age-dependent effects on eGFR, while the 11 stable-effect variants showed age-independent effects (Fig. 3a). The large extent of age-dependency for decline-associated variants was remarkable: near-zero effects on eGFR-levels among 40-year-old (even UMOD/PDILT; except PRAKG2), but large effects for 70-year-old individuals, much larger than cross-sectionally (e.g., for UMOD/PDILT rs77924615: −1.59 versus −0.90 mL/min/1.73 m2 per “bad” allele, respectively; for rs854922 near RRAGD: −0.55 versus −0.28; Supplementary Data 3). This suggests that age-dependent associations with eGFR become effective mainly around the age of 40 years, while stable associations are already effective before the age of 40 years and age-independent thereafter.
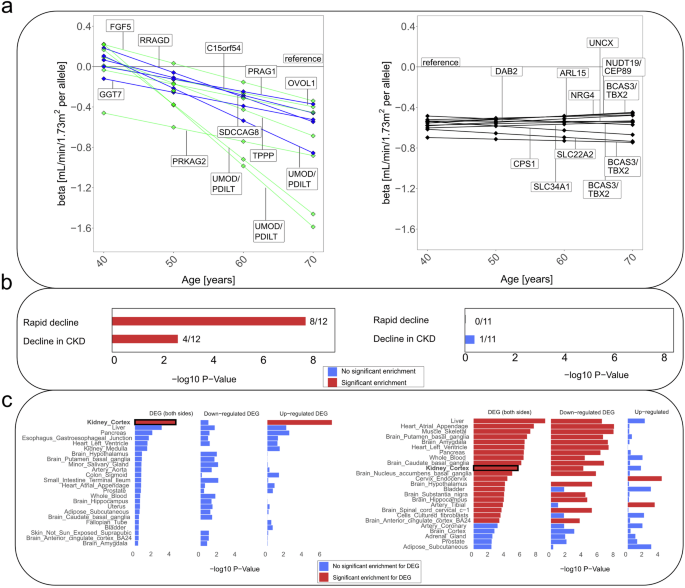
We contrasted the 12 decline-associated variants versus 11 stable-effect variants and underlying loci. a Shown are genetic effects on eGFR for 40-, 50-, 60-, 70-year-old individuals using LMM age model RI&RS 350K (beta derived as βmain + (age-50)*βdecline) for decline-associated variants (left; blue: novel, green: known) and stable-effect variants (right; black). Effect allele was the cross-sectionally eGFR-lowering allele23 (Supplementary Data 3). b We tested the 12 + 11 variants for association with two clinical progression traits using UKB 150K, rapid decline (ncases = 1211, ncontrols = 63,392, logistic regression) and decline in CKD (nCKD =13,116, mCKD = 116,944, LMM time model RI&RS; “Methods” section and Supplementary Table 6). Significant enrichment (Penrich < 0.05) of directionally consistent nominally significant associations was found among the 12 (left; 8/12, 4/12), but not among the 11 SNPs (right; 0/11, 1/11). c We evaluated genes in loci of the 12 + 11 variants regarding tissue-specific enrichment of differentially expressed genes (DEGs): shown are enrichment P values in decline-associated loci (left, among 256 genes) and stable-associated loci (right, among 182 genes; using FUMA, testing 54 tissue types, showing top 25; “Methods” section). Significant enrichment for DEGs (FDR < 0.05, red) was found for decline-associated loci only in kidney cortex (upregulated) and for stable-effect loci in various tissues (mostly downregulated, e.g., in liver, heart, muscle, pancreas, kidney cortex).
Robustness of findings regarding non-linear age effects and eGFR-variability
The approaches applied here and by others10,11,20,21 assume linearity in the global age effect on eGFR, the person-specific age effects on eGFR, and the age effect on the SNP-association with eGFR (i.e., modeling SNP-association with linear eGFR-decline). Allowing for non-linear relationships (adding quadratic terms; “Methods” section) did not alter results for the 12 + 11 SNP associations with linear eGFR-decline (Supplementary Fig. 6). Two variants, rs77924615 and rs13334589 in/around UMOD/PDILT, showed a small, but significant association with over-linear eGFR-decline (Supplementary Fig. 7; PSNPxage² < 0.05/23 = 2.2 × 10−3; Supplementary Data 4). Further analyses for these two variants pointed to 50 years as breakpoint for accelerated decline (Pbreakpoint50 = 6.3 × 10−56 and 1.7 × 10−5, respectively; Pbreakpoint40 = 0.45 and 0.50, Pbreakpoint60 = 0.04 and 0.04; “Methods” section).
Longitudinal data have also been used to test for SNP associations with trait variability15. When applying the model implemented in TrajGWAS15 (“Methods” section), all 12 decline-associated variants, but also 7 stable-effect variants were associated with eGFR-variability (P < 0.05/23 = 2.2 × 10−3; Supplementary Fig. 8). Thus, association with eGFR-variability answers a different question than association with eGFR-decline.
Decline-associated variants show SNP-by-age interaction in cross-sectional data
Decline-associated SNPs should show SNP-by-age interaction in cross-sectional data (UKB study-center baseline, n = 341,073; linear regression adjusted for sex, 20 principal components (PCs)): 10 of 12 showed PSNPxage < 0.05; when compared to effects on eGFR-decline in longitudinal data, interaction effects were similar (−0.010 to −0.048 mL/min/1.73 m2 per allele and year) and P values were larger, attributable to reduced power (Supplementary Data 5). None of the 11 stable-effect variants had PSNPxage < 0.05 with negative effect.
The cross-sectional data also gave us the opportunity to explore whether the age-dependency of the 12 SNP associations with eGFR was explained by their interaction with diabetes, HbA1c, hypertension, or systolic blood pressure (SBP). The SNP-by-age interaction effects remained the same when including SNP-by-diabetes, SNP-by-HbA1c, SNP-by-hypertension, or SNP-by-SBP interaction terms (Supplementary Fig. 9 and Supplementary Data 5).
Differential pattern of association with clinical progression traits between decline-associated versus stable-effect loci
From a clinical perspective, rapid eGFR-decline or eGFR-decline in CKD are of particular interest as surrogate for CKD progression3,27. Previous work on the genetics of these progression traits identified SNPs around UMOD/PDILT, PRKAG2, and TPPP11,20,21,28, suggesting an overlap with genetics of eGFR-decline in general population. We tested the 12 + 11 SNPs for association with rapid decline (ncases =1211, ncontrols = 63,392; “Methods” section) and with eGFR-decline in the subset of individuals with CKD (eGFR < 60 mL/min/1.73 m2, nckd = 13,116, mCKD = 116,944; “Methods” section). The 12 decline-associated variants were enriched for directionally consistent nominally significant association with rapid decline and eGFR-decline in CKD (Penrich =1.6 × 10−8 or 2.2 × 10−3, respectively), but the 11 stable-effect variants were not (Penrich = 1.0 or 0.43, respectively; Fig. 3b and Supplementary Table 6). Decline-associated variants contributing to these enrichments were near UMOD/PDILT (3), PRKAG2, and TPPP (confirmed for clinical progression traits), RRAGD, OVOL1, and C15orf54 (novel).
Both the 12 and 11 variants were enriched for association with the odds of having CKD (ncases =16,147, ncontrols = 332,128; Penrich = 2.4 × 10−16 and 9.8 × 10−11, respectively). Thus, decline-associated versus stable-effect variants showed a similar relevance for having/developing CKD, but a differential pattern for clinical progression traits.
Differential pattern of tissue-specific gene expression regulation in decline-associated versus stable-effect loci
We were interested in likely causal genes and potentially differential mechanisms implicated by the 12 decline-associated variants (10 loci) versus the 11 stable-effect variants (9 loci).
We annotated biological and statistical features to 256 and 182 genes in these loci (“Methods” section; Supplementary Data 6). We found accumulated evidence with ≥3 features for six genes to be likely causal for decline-associated loci (UMOD, PRKAG2, SDCCAG8, RRAGD, TPPP, FGF5) and for four genes for stable-effect loci (CPS1, SLC22A2, SLC34A1, UNCX; Table 4 and Supplementary Note 3). For the highlighted 6 + 4 = 10 genes, the locus index variant was in or very near (<25 kb) to the mapped gene and statistically highly likely the association-driving variant (22%–100% probability). Common-variant effects for Mendelian disease genes were found for both decline-associated and stable-effect variants; two genes known for a role in creatinine metabolism (creatinine production or tubular reuptake29,30) mapped to stable-effect loci.
While pathway-enrichment analyses were inconclusive (using Panther31,32, “Methods” section and Supplementary Note 3), analysis of tissue-specific enrichment for differentially expressed genes (DEGs) showed a strikingly differential pattern (using FUMA33, “Methods” section): significant enrichment for DEGs (false discovery rate, FDR < 0.05) was found only in kidney cortex for decline-associated loci (upregulated), yet in various tissues for stable-effect loci (mostly downregulated; e.g., in heart, liver, muscle, pancreas, kidney cortex; Fig. 3c). This suggests that decline-associated versus stable-effect loci differentiate kidney-specific versus cross-organ regulation of gene expression.
LMM-based longGWAS identifies five loci with genome-wide significance highlighting MUC1 for eGFR-decline
We now applied the LMM age model RI&RS in UKB 350K using the GMMAT/MAGEE34,35 implementation, which implements this model in a more efficient way than lme4 (“Methods” section). We tested the 595 variants and corroborated that association statistics for both implementations, GMMAT/MAGEE versus lme4, were identical (Supplementary Fig. 10 and Supplementary Data 7).
We used GMMAT/MAGEE to conduct a longGWAS, testing ~11 million autosomal variants (UK10K/HRC-imputed36, “Methods” section). We obtained results within 5 days (256 cores, 1 TB RAM) with little evidence for population stratification (lambda = 1.06).
We identified five loci associated with eGFR-decline at genome-wide significance (GC-corrected Pdecline < 5 × 10−8, “Methods” section, Fig. 4): the four loci already identified with Pdecline < 5 × 10−8 by the 595-search and one additional locus (MTX1/MUC1, novel for eGFR-decline compared to previous work10).
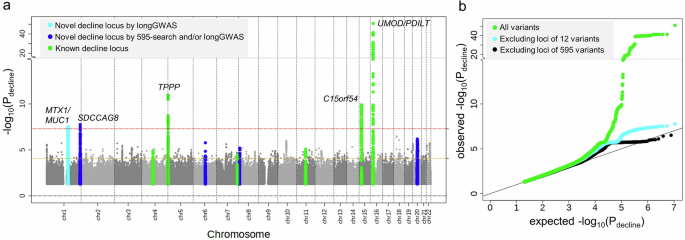
We conducted a genome-wide search for genetic variant association with eGFR-decline (Pdecline, GC-corrected, lambda = 1.06) using the LMM age model RI&RS 350K implemented in GMMAT/MAGEE34,35 (UKB 350K; n = 348,275, m = 1,520,382; testing 11 million SNPs with MAF ≥ 0.5%, imputation quality INFO ≥ 0.6). a Shown are association P values versus chromosomal position. We identified five loci at genome-wide significance (Pdecline < 5 × 10−8; red dashed horizontal line). Coloring highlights the overall 11 loci identified for eGFR-decline: 10 loci around the 12 variants identified by 595-search (Pdecline < 0.05/595 = 8.4 × 10−5, brown dashed horizontal line; 4 novel and 6 known for eGFR-decline in blue or green, respectively), and one novel locus for eGFR-decline now identified by longGWAS (cyan; lead variant rs2075570 in the 424 loci, but not among the 595 variants). Loci were derived by clumping based on variant position (d > 500kB between loci, “Methods” section). b Shown is the Quantile–Quantile (QQ) plot comparing the distribution of observed Pdecline with the distribution of Pdecline expected under the null hypothesis of “no association with eGFR-decline” (green: all variants; cyan: excluding the 10 loci around the 12 decline-associated variants; black: excluding the 424 loci around the 595 variants).
The lead variant of the MTX1/MUC1 locus, rs2075570 (Pdecline = 1.1 × 10−8), resided in the 424 loci known for cross-sectional eGFR, but was not among or correlated to the 595 variants (Pcross-sectional = 0.01 in Stanzick et al.23; Pcross-sectional = 0.80 in UKB; Supplementary Fig. 11a, b). Breakpoint analyses suggest a complex age-dependency of the rs2075570-association on eGFR (Supplementary Fig. 11c). rs2075570 modifies expression for MUC1 in tubolo-interstitial tissue37, (FDR < 5%), which suggests MUC1, a well-known gene for rare autosomal dominant tubulo-interstitial kidney disease38,39, as likely causal gene.
In total, we identified 13 independent variants (11 loci) for eGFR-decline: 7 variants (5 loci) with Pdecline < 5 × 10−8 by longGWAS and/or the 595-search and 6 variants (6 loci) by the 595-search (Pdecline < 0.05/595; Supplementary Table 7). LongGWAS results also enabled us to show full regional association signals for decline-associated loci, which align well with respective signals from cross-sectional analyses (Supplementary Fig. 12), except for the MTX/MUC1 signal (Supplementary Fig. 11a).
link